Nanographics has published a scientific paper presented at the IEEE VIS 2019 conference. The paper presents a novel technique to procedurally animate complex molecular structures based on measured data.
Abstract
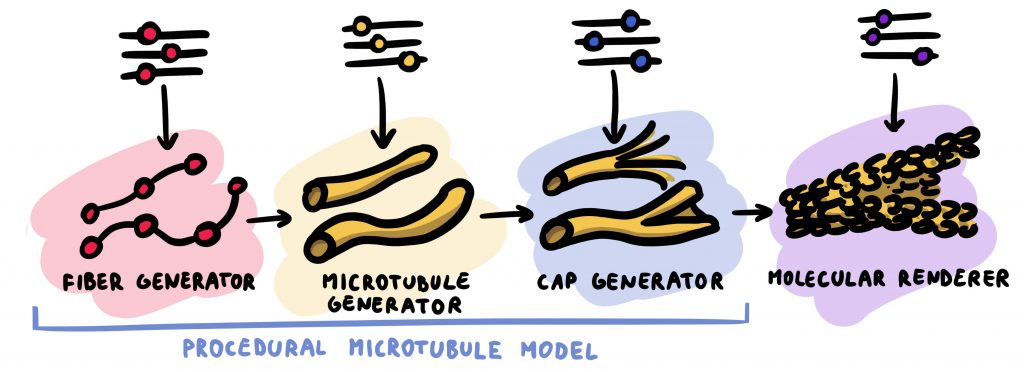
Biologists often use computer graphics to visualize structures, which due to physical limitations are not possible to image with a microscope. One example for such structures are microtubules, which are present in every eukaryotic cell. They are part of the cytoskeleton maintaining the shape of the cell and playing a key role in the cell division. In this paper, we propose a scientifically-accurate multi-scale procedural model of microtubule dynamics as a novel application scenario for procedural animation, which can generate visualizations of their overall shape, molecular structure, as well as animations of the dynamic behaviour of their growth and disassembly. The model is spanning from tens of micrometers down to atomic resolution. All the aspects of the model are driven by scientific data. The advantage over a traditional, manual animation approach is that when the underlying data change, for instance due to new evidence, the model can be recreated immediately. The procedural animation concept is presented in its generic form, with several novel extensions, facilitating an easy translation to other domains with emergent multi-scale behavior.
Paper: “Multi-Scale Procedural Animations of Microtubule Dynamics Based on Measured Data”